 The dNTPs formed from this activity would reduce our concentration of radiolabeled dNTP's, and cause gaps of unlabeled sequences in our new copy. The reaction products with 30mer primers are B1 (Klenow fragment using the. If DNA pol I was used in this example, it could degrade (5'-3' exnuclease) some of these strands (ones that did not have a primer attached). polymerase was not useful for low temperature cycle primer extension. The only reasoning I could come up with was, with the use of denaturing dsDNA, there would be free floating ssDNA. However, when it comes to primer extension radio labeling (using a single strand as the template - cDNA, or maybe even denaturing of dsDNA?), the klenow fragment is used.why? It seems when it comes to nick translation radiolabeling, DNA pol I is used understandable since the 5'-3' exonuclease is needed to degrade the non-template strand, allowing for the radiolabeling. Random primers (octamers) are annealed to the denatured DNA template and extended by Klenow fragment in the presence of biotin. 
My question revolves around radioactive labeling and its applications. Degrade 3' over hangs (again it seems DNA pol I could do this, or would the 5'-3' exonuclease begin to degrade the opposite strand?) The enzyme exhibits DNA synthesis and proofreading (35) nuclease activities and, in the absence of the holoenzyme’s (53) nuclease domain, displays a moderate strand displacement activity during DNA synthesis. coli Polymerase I DNA-dependent repair enzyme. Fill in 5' over hangs (Can't DNA pol I do this?) Protease-mediated allele-specific extension (PrASE) is a flexible assay and can be performed at lower temperatures using thermolabile enzymes, such as the. Product Details Klenow Fragment is a mesophilic DNA polymerase derived from the E. Many sources say klenow fragments are able to synthesize dsDNA from cDNA (make dsDNA from a single strand) but can't DNA pol I do this? (of course both needing primers).Īdditionally, it is pointed out that Klenow fragments can be used to form blunt ends: 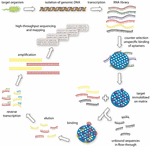
from publication: Photoresponsive nucleic acids for. This allows routine incorporation of Radiolabeled nucleotide at greater than 60, in less than 30-60 minutes, providing DNA probe specific activities of greater than 109 dpm/g DNA. Download scientific diagram Strategy for photoregulating primer extension by the Klenow fragment (KF). My question though stems from many of the sources I've been scouring. Because Klenow fragment lacks the 5’ 3’ exonuclease activity, the use of Klenow fragment in primer extension avoids the loss of incorporated label. Since it is provided in its inactive form, it can be added to a reaction without the fear of pre-PCR misprimed primers being extended. Some more information regarding my question:įrom my understanding, the only difference that a klenow fragment has over DNA polymerase I, is its removal of 5'-3' exonuclease activity (additionally, removal of 3'-5' exonuclease activity if specified). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
